Bayesian inference for NLMMs with group-specific coefficients that have unknown covariance matrices with flexible priors.
Usage
stan_nlmer(
formula,
data = NULL,
subset,
weights,
na.action,
offset,
contrasts = NULL,
...,
prior = normal(autoscale = TRUE),
prior_aux = exponential(autoscale = TRUE),
prior_covariance = decov(),
prior_PD = FALSE,
algorithm = c("sampling", "meanfield", "fullrank"),
adapt_delta = NULL,
QR = FALSE,
sparse = FALSE
)Arguments
- formula, data
Same as for
nlmer. We strongly advise against omitting thedataargument. Unlessdatais specified (and is a data frame) many post-estimation functions (includingupdate,loo,kfold) are not guaranteed to work properly.- subset, weights, offset
Same as
glm.- na.action, contrasts
Same as
glm, but rarely specified.- ...
Further arguments passed to the function in the rstan package (
sampling,vb, oroptimizing), corresponding to the estimation method named byalgorithm. For example, ifalgorithmis"sampling"it is possible to specifyiter,chains,cores, and other MCMC controls.Another useful argument that can be passed to rstan via
...isrefresh, which specifies how often to print updates when sampling (i.e., show the progress everyrefreshiterations).refresh=0turns off the iteration updates.- prior
The prior distribution for the (non-hierarchical) regression coefficients.
The default priors are described in the vignette Prior Distributions for rstanarm Models. If not using the default,
priorshould be a call to one of the various functions provided by rstanarm for specifying priors. The subset of these functions that can be used for the prior on the coefficients can be grouped into several "families":Family Functions Student t family normal,student_t,cauchyHierarchical shrinkage family hs,hs_plusLaplace family laplace,lassoProduct normal family product_normalSee the priors help page for details on the families and how to specify the arguments for all of the functions in the table above. To omit a prior —i.e., to use a flat (improper) uniform prior—
priorcan be set toNULL, although this is rarely a good idea.Note: Unless
QR=TRUE, ifprioris from the Student t family or Laplace family, and if theautoscaleargument to the function used to specify the prior (e.g.normal) is left at its default and recommended value ofTRUE, then the default or user-specified prior scale(s) may be adjusted internally based on the scales of the predictors. See the priors help page and the Prior Distributions vignette for details on the rescaling and theprior_summaryfunction for a summary of the priors used for a particular model.- prior_aux
The prior distribution for the "auxiliary" parameter (if applicable). The "auxiliary" parameter refers to a different parameter depending on the
family. For Gaussian modelsprior_auxcontrols"sigma", the error standard deviation. For negative binomial modelsprior_auxcontrols"reciprocal_dispersion", which is similar to the"size"parameter ofrnbinom: smaller values of"reciprocal_dispersion"correspond to greater dispersion. For gamma modelsprior_auxsets the prior on to the"shape"parameter (see e.g.,rgamma), and for inverse-Gaussian models it is the so-called"lambda"parameter (which is essentially the reciprocal of a scale parameter). Binomial and Poisson models do not have auxiliary parameters.The default prior is described in the vignette Prior Distributions for rstanarm Models. If not using the default,
prior_auxcan be a call toexponentialto use an exponential distribution, ornormal,student_torcauchy, which results in a half-normal, half-t, or half-Cauchy prior. Seepriorsfor details on these functions. To omit a prior —i.e., to use a flat (improper) uniform prior— setprior_auxtoNULL.- prior_covariance
Cannot be
NULL; seedecovfor more information about the default arguments.- prior_PD
A logical scalar (defaulting to
FALSE) indicating whether to draw from the prior predictive distribution instead of conditioning on the outcome.- algorithm
A string (possibly abbreviated) indicating the estimation approach to use. Can be
"sampling"for MCMC (the default),"optimizing"for optimization,"meanfield"for variational inference with independent normal distributions, or"fullrank"for variational inference with a multivariate normal distribution. Seerstanarm-packagefor more details on the estimation algorithms. NOTE: not all fitting functions support all four algorithms.- adapt_delta
Only relevant if
algorithm="sampling". See the adapt_delta help page for details.- QR
A logical scalar defaulting to
FALSE, but ifTRUEapplies a scaledqrdecomposition to the design matrix. The transformation does not change the likelihood of the data but is recommended for computational reasons when there are multiple predictors. See the QR-argument documentation page for details on how rstanarm does the transformation and important information about how to interpret the prior distributions of the model parameters when usingQR=TRUE.- sparse
A logical scalar (defaulting to
FALSE) indicating whether to use a sparse representation of the design (X) matrix. IfTRUE, the the design matrix is not centered (since that would destroy the sparsity) and likewise it is not possible to specify bothQR = TRUEandsparse = TRUE. Depending on how many zeros there are in the design matrix, settingsparse = TRUEmay make the code run faster and can consume much less RAM.
Value
A stanreg object is returned
for stan_nlmer.
Details
The stan_nlmer function is similar in syntax to
nlmer but rather than performing (approximate) maximum
marginal likelihood estimation, Bayesian estimation is by default performed
via MCMC. The Bayesian model adds independent priors on the "coefficients"
— which are really intercepts — in the same way as
stan_nlmer and priors on the terms of a decomposition of the
covariance matrices of the group-specific parameters. See
priors for more information about the priors.
The supported transformation functions are limited to the named
"self-starting" functions in the stats library:
SSasymp, SSasympOff,
SSasympOrig, SSbiexp,
SSfol, SSfpl,
SSgompertz, SSlogis,
SSmicmen, and SSweibull.
See also
stanreg-methods and
nlmer.
The vignette for stan_glmer, which also discusses
stan_nlmer models. https://mc-stan.org/rstanarm/articles/
Examples
if (.Platform$OS.type != "windows" || .Platform$r_arch !="i386") {
# \donttest{
data("Orange", package = "datasets")
Orange$circumference <- Orange$circumference / 100
Orange$age <- Orange$age / 100
fit <- stan_nlmer(
circumference ~ SSlogis(age, Asym, xmid, scal) ~ Asym|Tree,
data = Orange,
# for speed only
chains = 1,
iter = 1000
)
print(fit)
posterior_interval(fit)
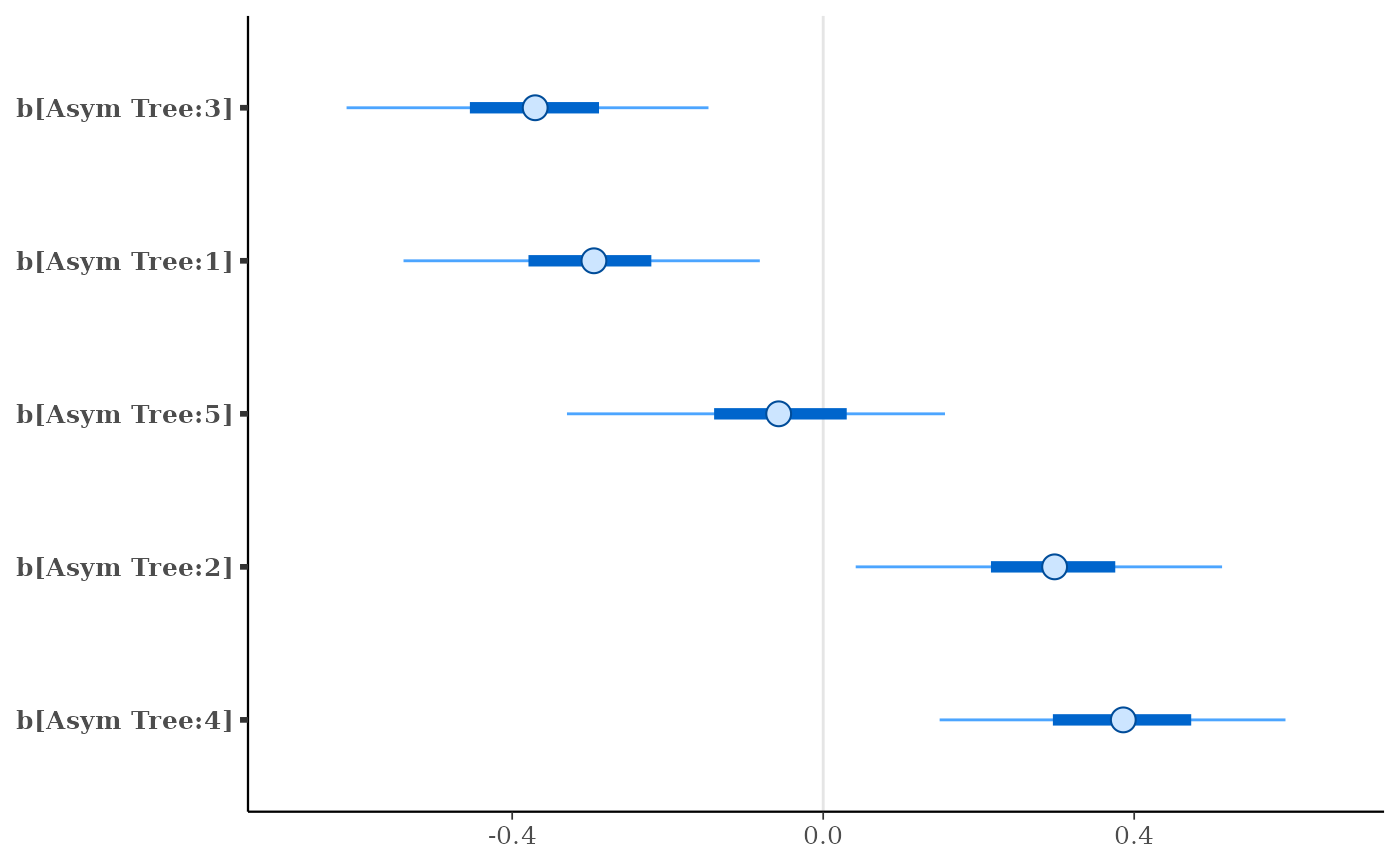
plot(fit, regex_pars = "b\\[")
# }
}
#> Warning: number of iterations exceeded maximum of 0
#>
#> SAMPLING FOR MODEL 'continuous' NOW (CHAIN 1).
#> Chain 1:
#> Chain 1: Gradient evaluation took 4e-05 seconds
#> Chain 1: 1000 transitions using 10 leapfrog steps per transition would take 0.4 seconds.
#> Chain 1: Adjust your expectations accordingly!
#> Chain 1:
#> Chain 1:
#> Chain 1: Iteration: 1 / 1000 [ 0%] (Warmup)
#> Chain 1: Iteration: 100 / 1000 [ 10%] (Warmup)
#> Chain 1: Iteration: 200 / 1000 [ 20%] (Warmup)
#> Chain 1: Iteration: 300 / 1000 [ 30%] (Warmup)
#> Chain 1: Iteration: 400 / 1000 [ 40%] (Warmup)
#> Chain 1: Iteration: 500 / 1000 [ 50%] (Warmup)
#> Chain 1: Iteration: 501 / 1000 [ 50%] (Sampling)
#> Chain 1: Iteration: 600 / 1000 [ 60%] (Sampling)
#> Chain 1: Iteration: 700 / 1000 [ 70%] (Sampling)
#> Chain 1: Iteration: 800 / 1000 [ 80%] (Sampling)
#> Chain 1: Iteration: 900 / 1000 [ 90%] (Sampling)
#> Chain 1: Iteration: 1000 / 1000 [100%] (Sampling)
#> Chain 1:
#> Chain 1: Elapsed Time: 0.764 seconds (Warm-up)
#> Chain 1: 0.383 seconds (Sampling)
#> Chain 1: 1.147 seconds (Total)
#> Chain 1:
#> stan_nlmer
#> family: gaussian [inv_SSlogis]
#> formula: circumference ~ SSlogis(age, Asym, xmid, scal) ~ Asym | Tree
#> observations: 35
#> ------
#> Median MAD_SD
#> Asym 1.9 0.1
#> xmid 7.2 0.3
#> scal 3.4 0.3
#>
#> Auxiliary parameter(s):
#> Median MAD_SD
#> sigma 0.1 0.0
#>
#> Error terms:
#> Groups Name Std.Dev.
#> Tree Asym 0.313
#> Residual 0.089
#> Num. levels: Tree 5
#>
#> ------
#> * For help interpreting the printed output see ?print.stanreg
#> * For info on the priors used see ?prior_summary.stanreg