Medians and central interval estimates of posterior or prior predictive
distributions. Each of these functions makes the same plot as the
corresponding ppc_ function but without plotting any
observed data y. The Plot Descriptions section at PPC-intervals has
details on the individual plots.
Usage
ppd_intervals(
ypred,
x = NULL,
...,
prob = 0.5,
prob_outer = 0.9,
alpha = 0.33,
size = 1,
fatten = 2.5,
linewidth = 1
)
ppd_intervals_grouped(
ypred,
x = NULL,
group,
...,
facet_args = list(),
prob = 0.5,
prob_outer = 0.9,
alpha = 0.33,
size = 1,
fatten = 2.5,
linewidth = 1
)
ppd_ribbon(
ypred,
x = NULL,
...,
prob = 0.5,
prob_outer = 0.9,
alpha = 0.33,
size = 0.25
)
ppd_ribbon_grouped(
ypred,
x = NULL,
group,
...,
facet_args = list(),
prob = 0.5,
prob_outer = 0.9,
alpha = 0.33,
size = 0.25
)
ppd_intervals_data(
ypred,
x = NULL,
group = NULL,
...,
prob = 0.5,
prob_outer = 0.9
)
ppd_ribbon_data(
ypred,
x = NULL,
group = NULL,
...,
prob = 0.5,
prob_outer = 0.9
)Arguments
- ypred
An
SbyNmatrix of draws from the posterior (or prior) predictive distribution, or aposterior::drawsobject. The number of rows,S, is the size of the posterior (or prior) sample used to generateypred. The number of columns,N, is the number of predicted observations.- x
A numeric vector to use as the x-axis variable. For example,
xcould be a predictor variable from a regression model, a time variable for time-series models, etc. Ifxis missing orNULLthen the observation index is used for the x-axis.- ...
Currently unused.
- prob, prob_outer
Values between
0and1indicating the desired probability mass to include in the inner and outer intervals. The defaults areprob=0.5andprob_outer=0.9.- alpha, size, fatten, linewidth
Arguments passed to geoms. For ribbon plots
alphais passed toggplot2::geom_ribbon()to control the opacity of the outer ribbon andsizeis passed toggplot2::geom_line()to control the size of the line representing the median prediction (size=0will remove the line). For interval plotsalpha,size,fatten, andlinewidthare passed toggplot2::geom_pointrange()(fatten=0will remove the point estimates).- group
A grouping variable of the same length as
y. Will be coerced to factor if not already a factor. Each value ingroupis interpreted as the group level pertaining to the corresponding observation.- facet_args
A named list of arguments (other than
facets) passed toggplot2::facet_wrap()orggplot2::facet_grid()to control faceting. Note: ifscalesis not included infacet_argsthen bayesplot may usescales="free"as the default (depending on the plot) instead of the ggplot2 default ofscales="fixed".
Value
The plotting functions return a ggplot object that can be further
customized using the ggplot2 package. The functions with suffix
_data() return the data that would have been drawn by the plotting
function.
References
Gabry, J. , Simpson, D. , Vehtari, A. , Betancourt, M. and Gelman, A. (2019), Visualization in Bayesian workflow. J. R. Stat. Soc. A, 182: 389-402. doi:10.1111/rssa.12378. (journal version, arXiv preprint, code on GitHub)
See also
Other PPDs:
PPD-distributions,
PPD-overview,
PPD-test-statistics
Examples
color_scheme_set("brightblue")
ypred <- example_yrep_draws()
x <- example_x_data()
group <- example_group_data()
ppd_intervals(ypred[, 1:50])
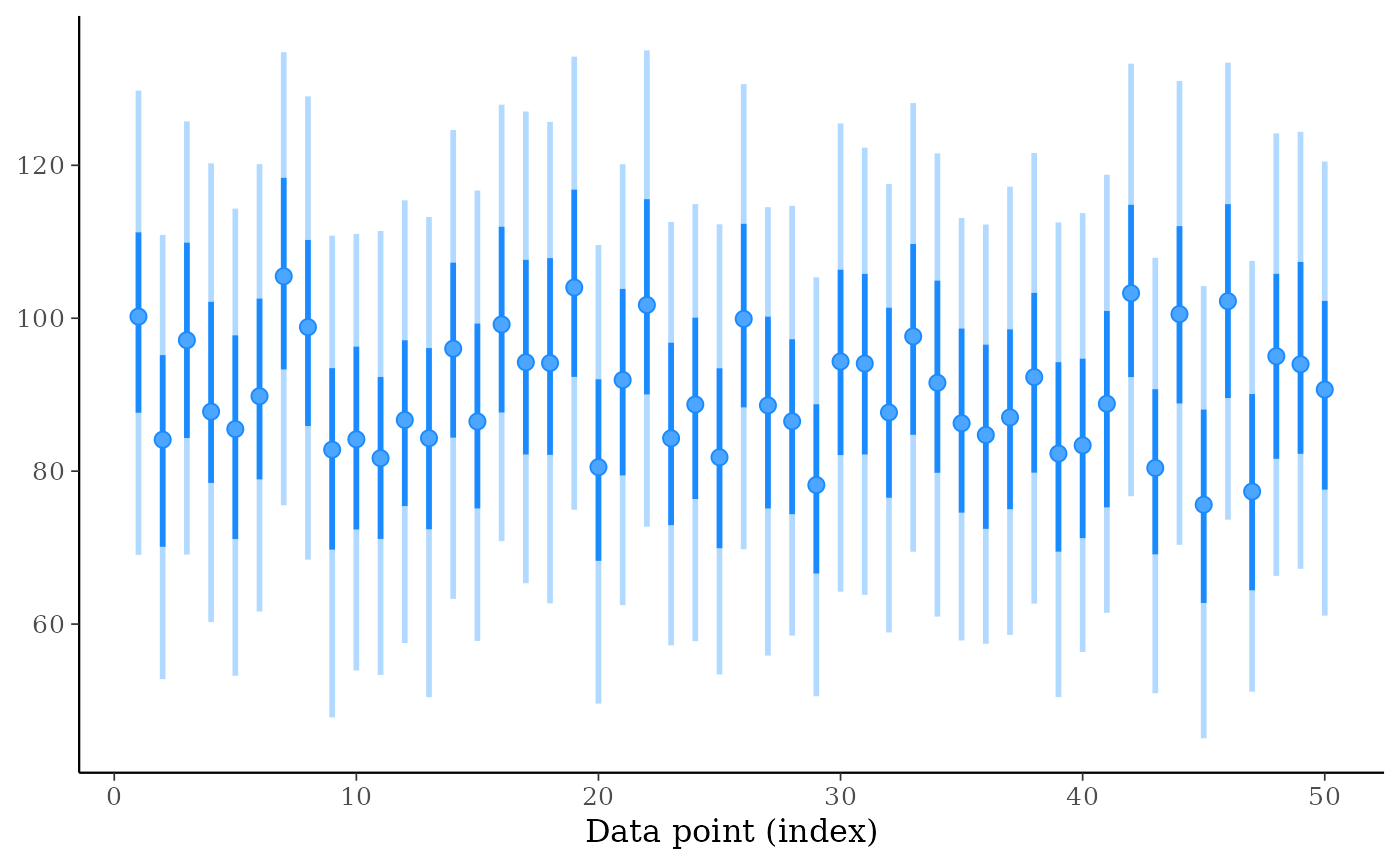 ppd_intervals(ypred[, 1:50], fatten = 0)
ppd_intervals(ypred[, 1:50], fatten = 0)
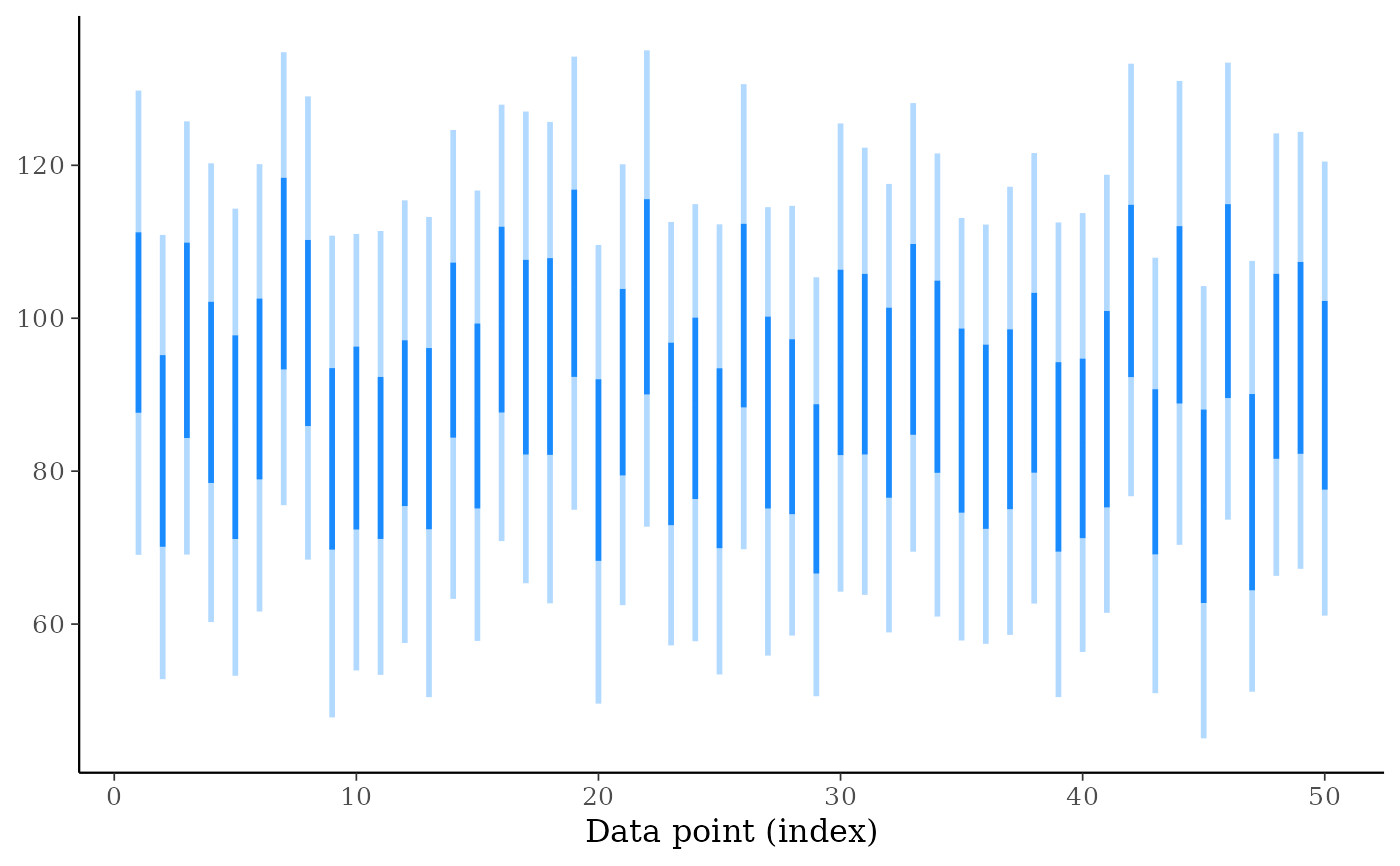 ppd_intervals(ypred[, 1:50], fatten = 0, linewidth = 2)
ppd_intervals(ypred[, 1:50], fatten = 0, linewidth = 2)
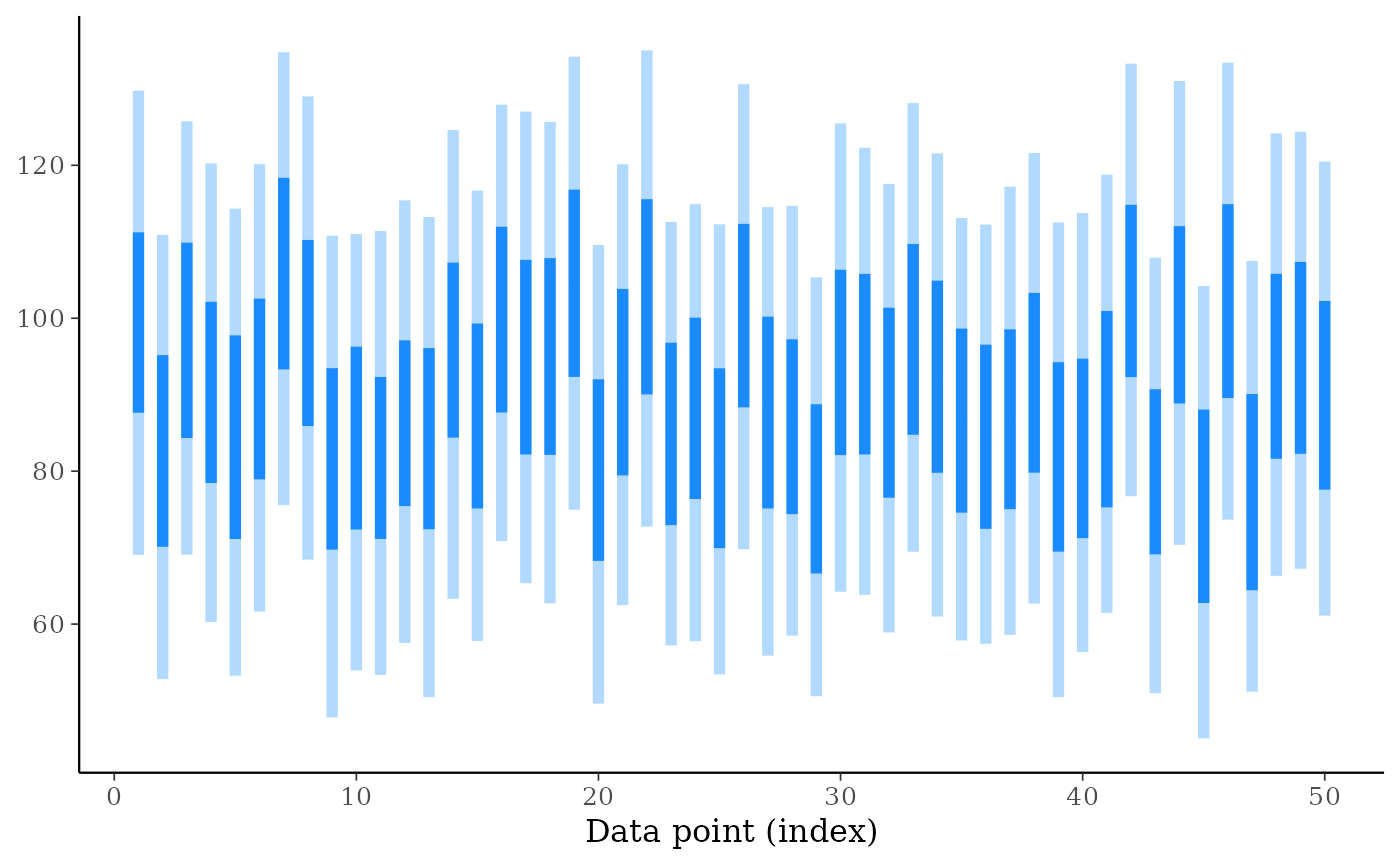 ppd_intervals(ypred[, 1:50], prob_outer = 0.75, fatten = 0, linewidth = 2)
ppd_intervals(ypred[, 1:50], prob_outer = 0.75, fatten = 0, linewidth = 2)
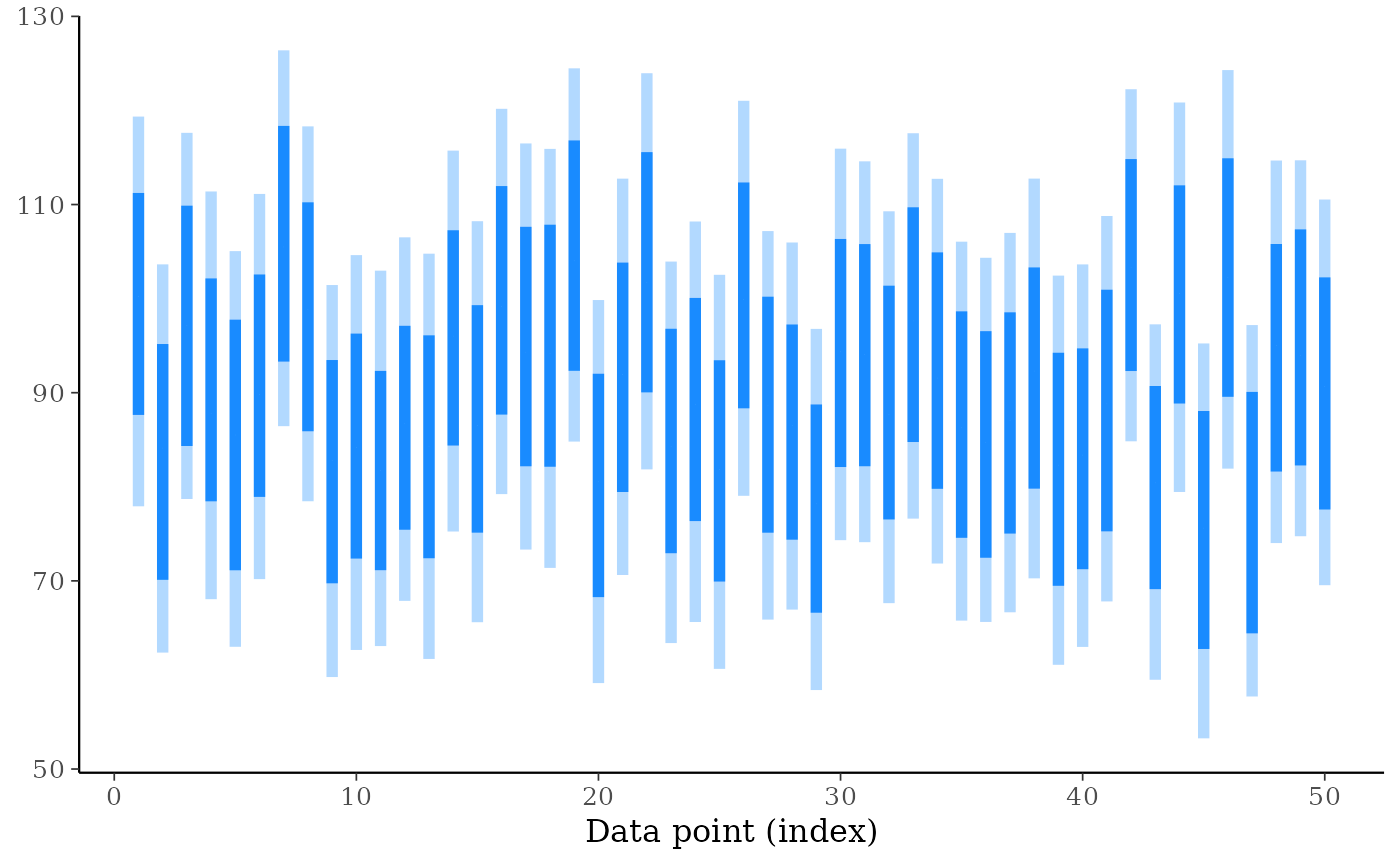 # put a predictor variable on the x-axis
ppd_intervals(ypred[, 1:100], x = x[1:100], fatten = 1) +
ggplot2::labs(y = "Prediction", x = "Some variable of interest")
# put a predictor variable on the x-axis
ppd_intervals(ypred[, 1:100], x = x[1:100], fatten = 1) +
ggplot2::labs(y = "Prediction", x = "Some variable of interest")
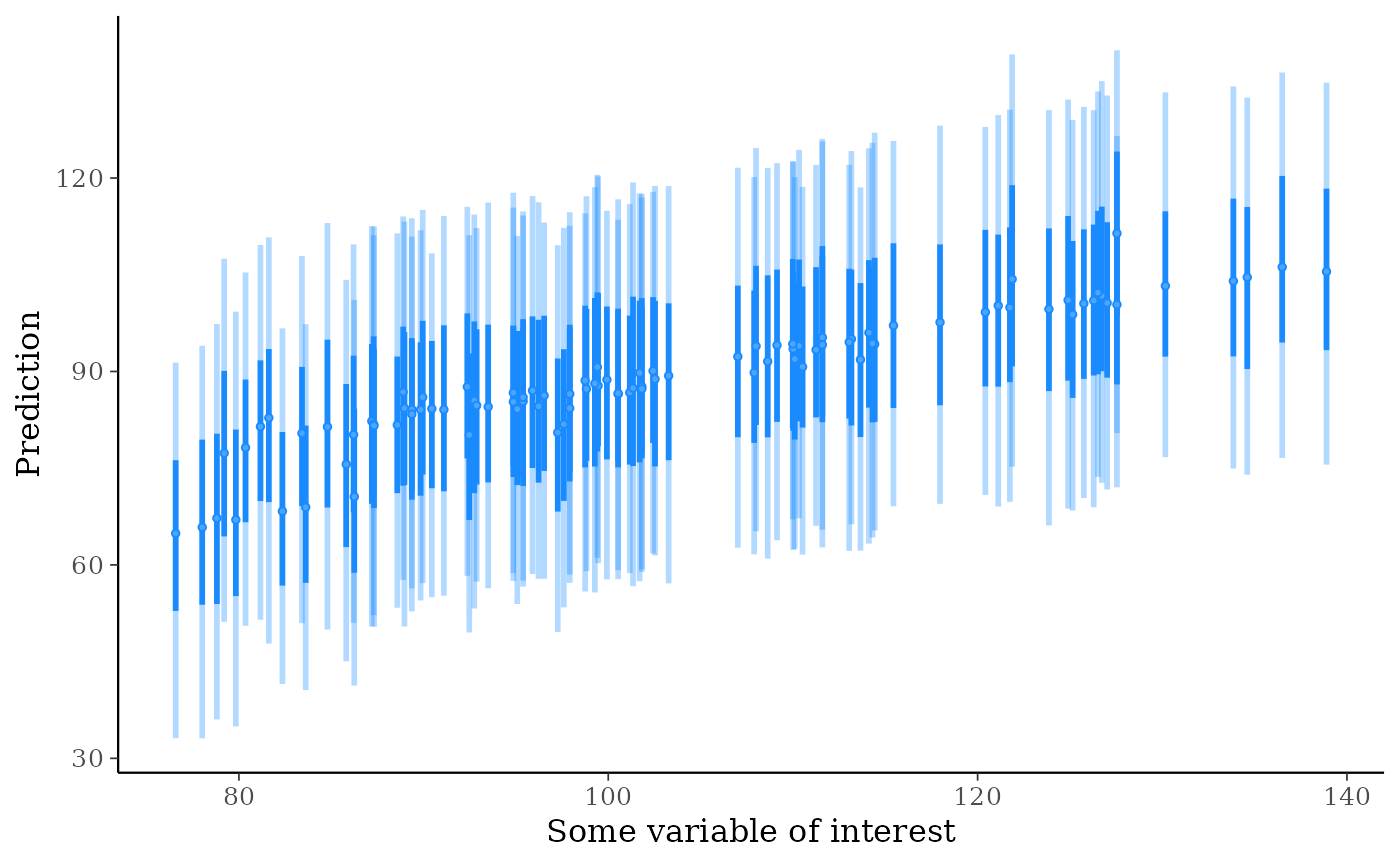 # with a grouping variable too
ppd_intervals_grouped(
ypred = ypred[, 1:100],
x = x[1:100],
group = group[1:100],
size = 2,
fatten = 0,
facet_args = list(nrow = 2)
)
# with a grouping variable too
ppd_intervals_grouped(
ypred = ypred[, 1:100],
x = x[1:100],
group = group[1:100],
size = 2,
fatten = 0,
facet_args = list(nrow = 2)
)
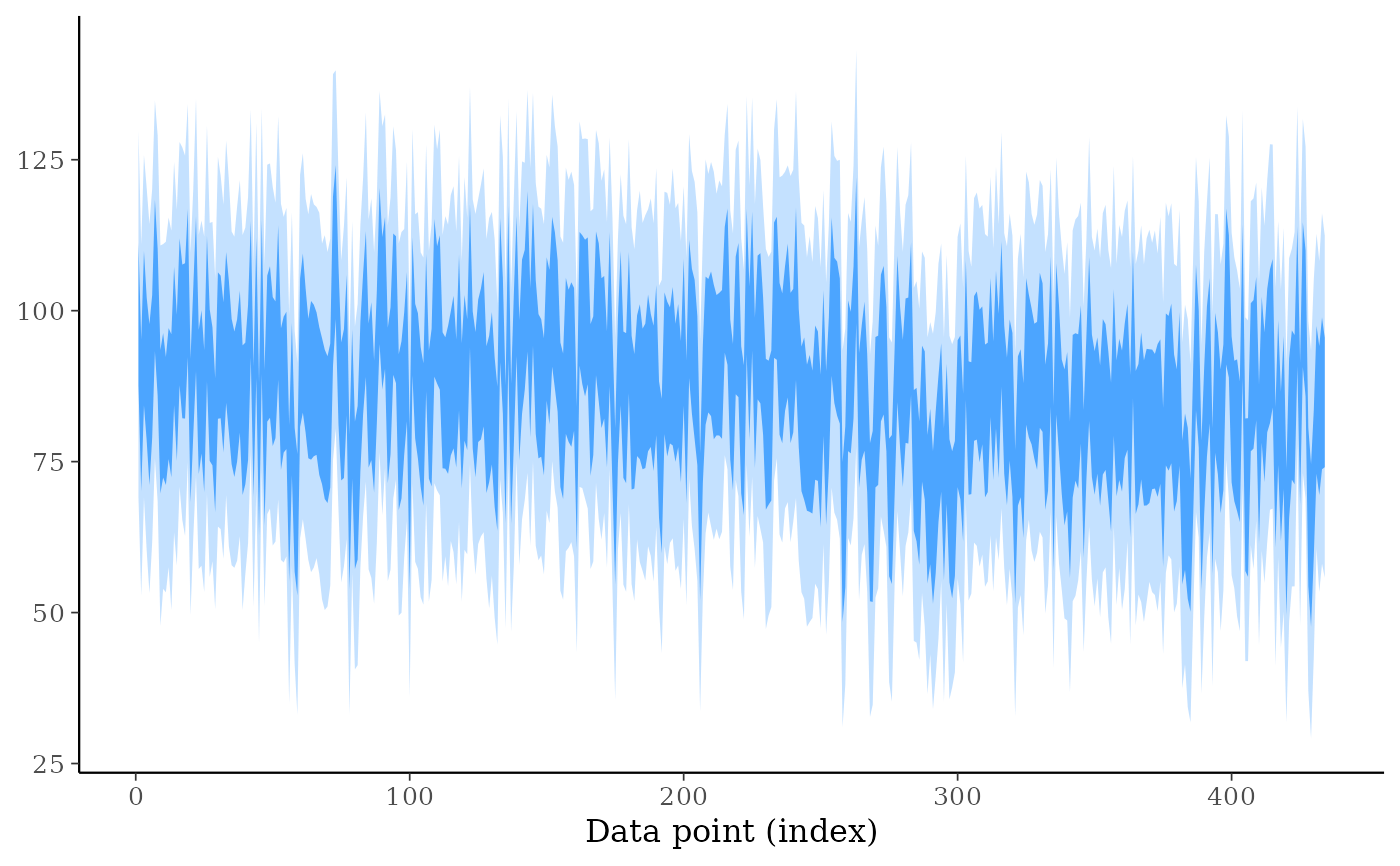
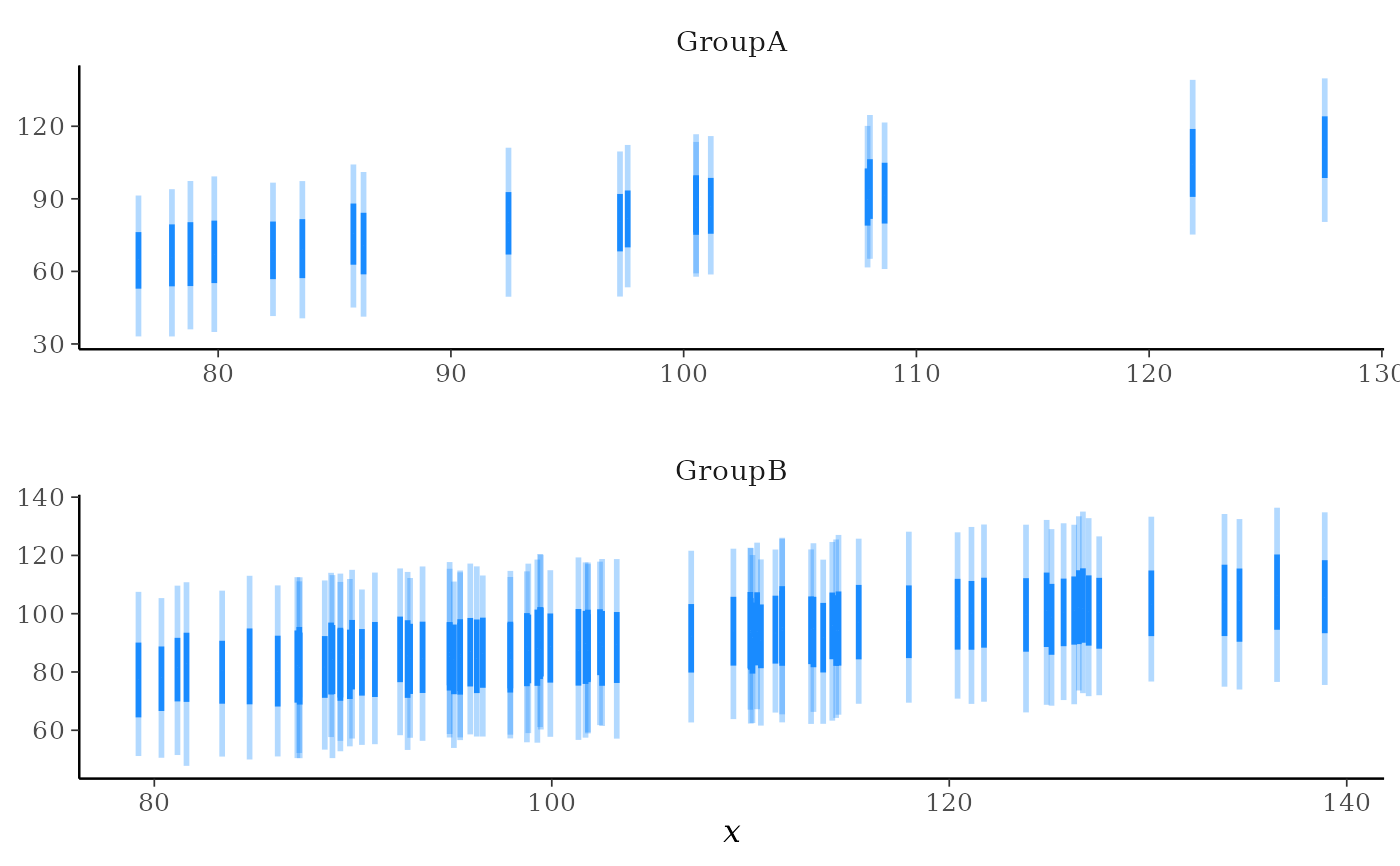 # even reducing size, ppd_intervals is too cluttered when there are many
# observations included (ppd_ribbon is better)
ppd_intervals(ypred, size = 0.5, fatten = 0.1, linewidth = 0.5)
# even reducing size, ppd_intervals is too cluttered when there are many
# observations included (ppd_ribbon is better)
ppd_intervals(ypred, size = 0.5, fatten = 0.1, linewidth = 0.5)
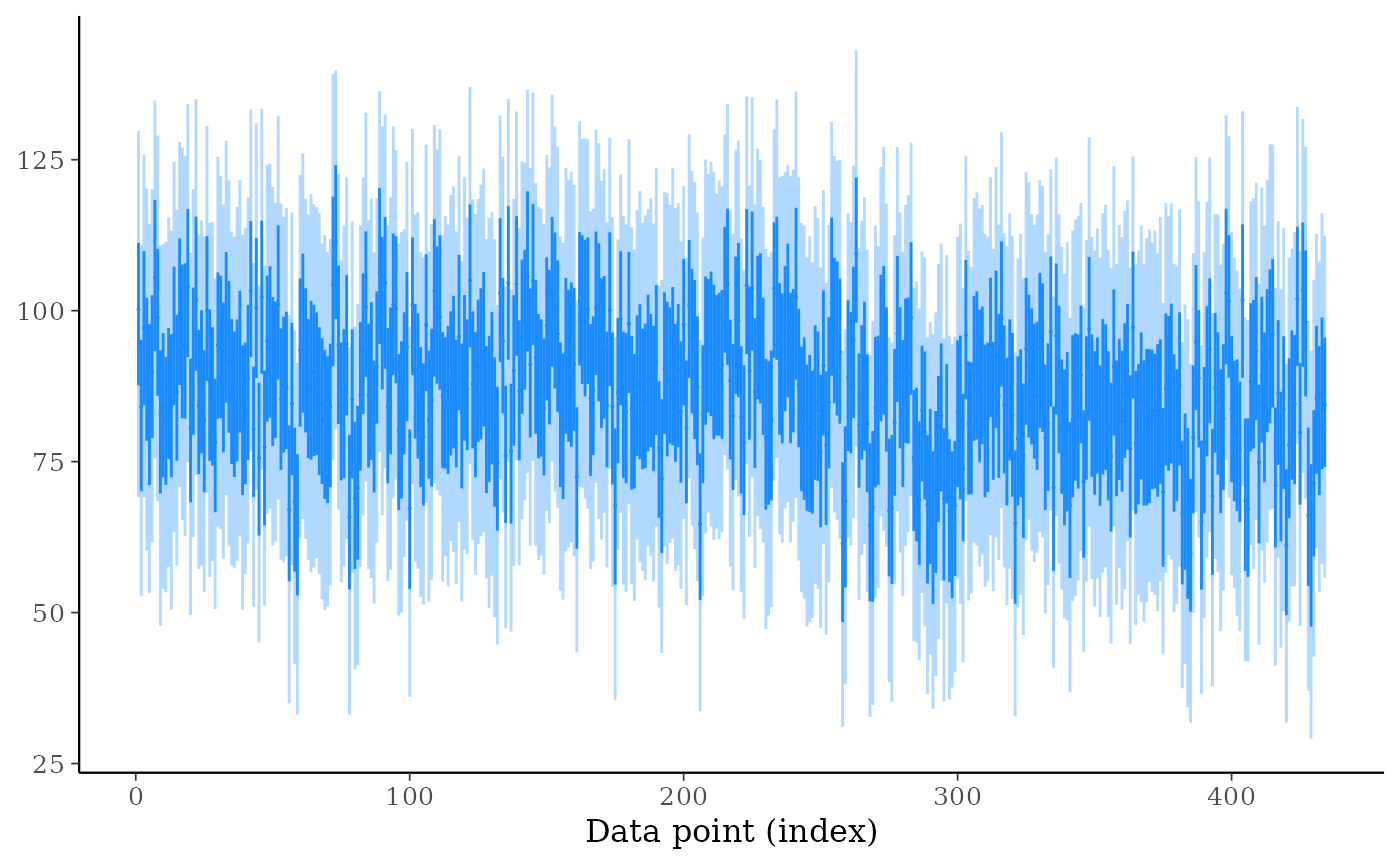 ppd_ribbon(ypred)
ppd_ribbon(ypred)
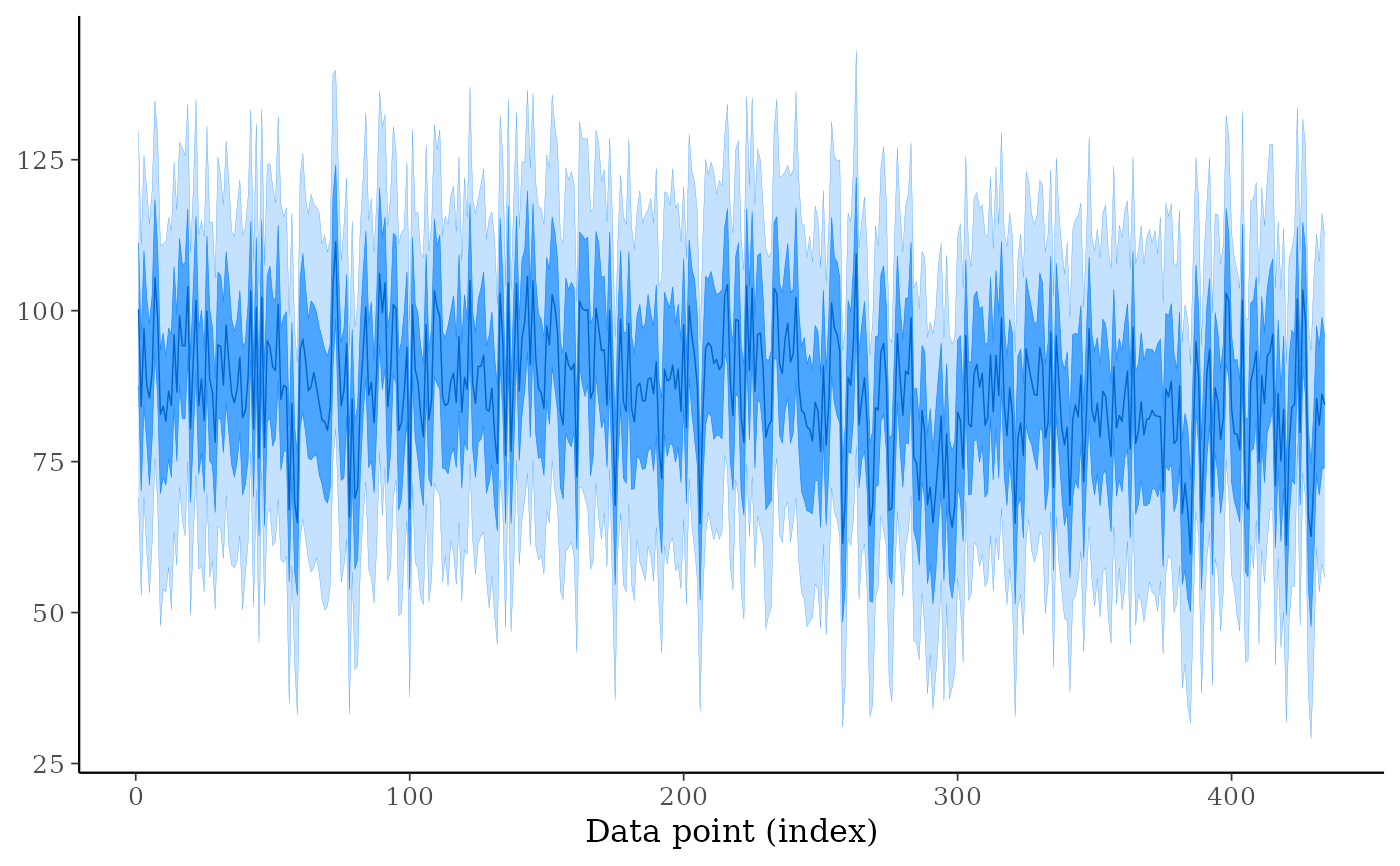 ppd_ribbon(ypred, size = 0) # remove line showing median prediction
ppd_ribbon(ypred, size = 0) # remove line showing median prediction