Compute a Bayesian version of R-squared or LOO-adjusted R-squared for regression models.
Source:R/bayes_R2.R
bayes_R2.stanreg.RdCompute a Bayesian version of R-squared or LOO-adjusted R-squared for regression models.
Usage
# S3 method for class 'stanreg'
bayes_R2(object, ..., re.form = NULL)
# S3 method for class 'stanreg'
loo_R2(object, ...)Arguments
- object
A fitted model object returned by one of the rstanarm modeling functions. See
stanreg-objects.- ...
Currently ignored.
- re.form
For models with group-level terms,
re.formis passed toposterior_epredif specified.
Value
A vector of R-squared values with length equal to the posterior sample size (the posterior distribution of R-squared).
References
Andrew Gelman, Ben Goodrich, Jonah Gabry, and Aki Vehtari (2019). R-squared for Bayesian regression models. The American Statistician, to appear. doi:10.1080/00031305.2018.1549100 (Article, Notebook)
Examples
if (.Platform$OS.type != "windows" || .Platform$r_arch != "i386") {
fit <- stan_glm(
mpg ~ wt + cyl,
data = mtcars,
QR = TRUE,
chains = 2,
refresh = 0
)
rsq <- bayes_R2(fit)
print(median(rsq))
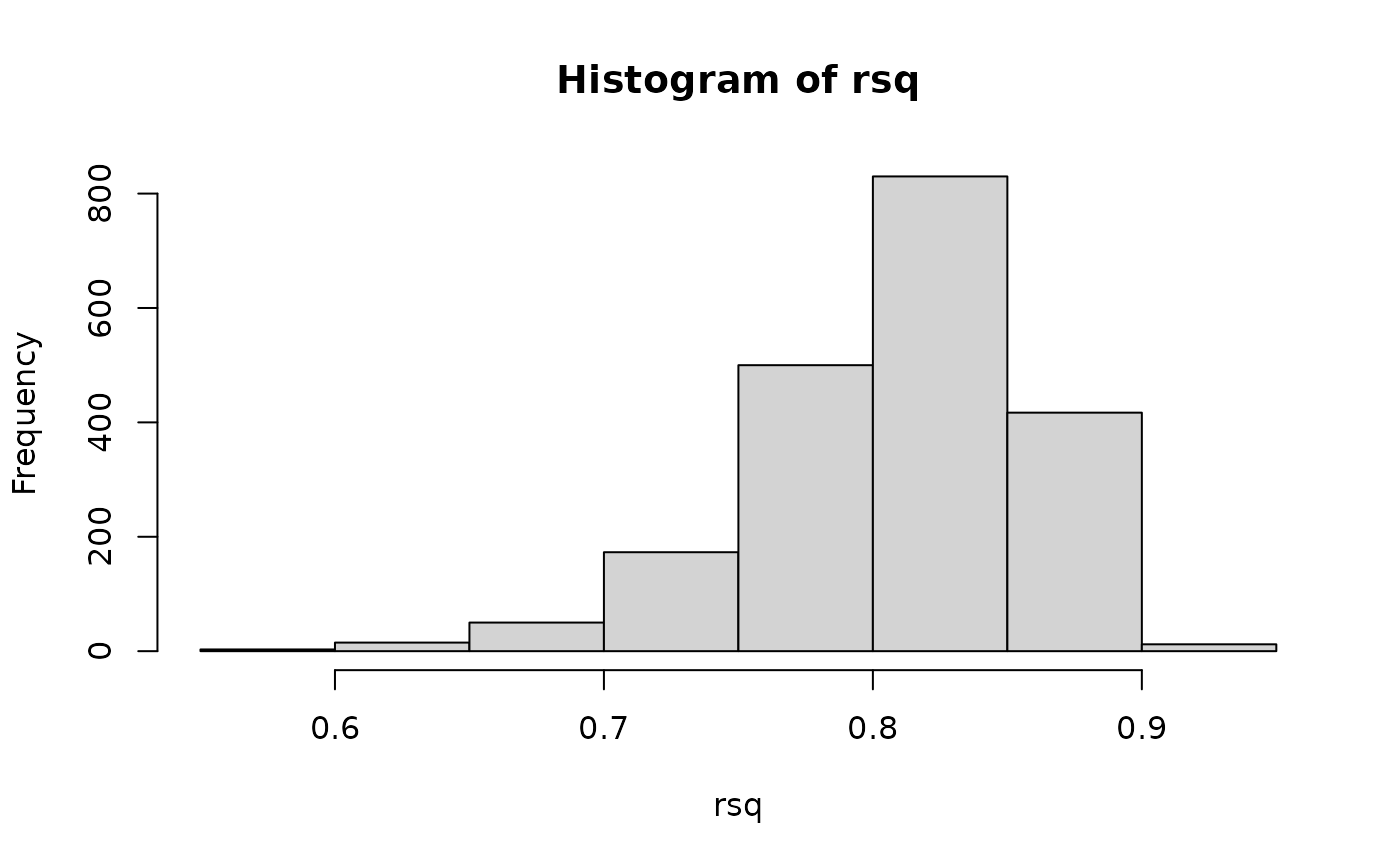
hist(rsq)
loo_rsq <- loo_R2(fit)
print(median(loo_rsq))
# multilevel binomial model
if (!exists("example_model")) example(example_model)
print(example_model)
median(bayes_R2(example_model))
median(bayes_R2(example_model, re.form = NA)) # exclude group-level
}
#> [1] 0.8156549
 #> [1] 0.7993705
#> stan_glmer
#> family: binomial [logit]
#> formula: cbind(incidence, size - incidence) ~ size + period + (1 | herd)
#> observations: 56
#> ------
#> Median MAD_SD
#> (Intercept) -1.5 0.6
#> size 0.0 0.0
#> period2 -1.0 0.3
#> period3 -1.1 0.4
#> period4 -1.6 0.5
#>
#> Error terms:
#> Groups Name Std.Dev.
#> herd (Intercept) 0.76
#> Num. levels: herd 15
#>
#> ------
#> * For help interpreting the printed output see ?print.stanreg
#> * For info on the priors used see ?prior_summary.stanreg
#> [1] 0.6206511
#> [1] 0.7993705
#> stan_glmer
#> family: binomial [logit]
#> formula: cbind(incidence, size - incidence) ~ size + period + (1 | herd)
#> observations: 56
#> ------
#> Median MAD_SD
#> (Intercept) -1.5 0.6
#> size 0.0 0.0
#> period2 -1.0 0.3
#> period3 -1.1 0.4
#> period4 -1.6 0.5
#>
#> Error terms:
#> Groups Name Std.Dev.
#> herd (Intercept) 0.76
#> Num. levels: herd 15
#>
#> ------
#> * For help interpreting the printed output see ?print.stanreg
#> * For info on the priors used see ?prior_summary.stanreg
#> [1] 0.6206511